|
10/13/2023 0 Comments Klenow fragment mutagenesis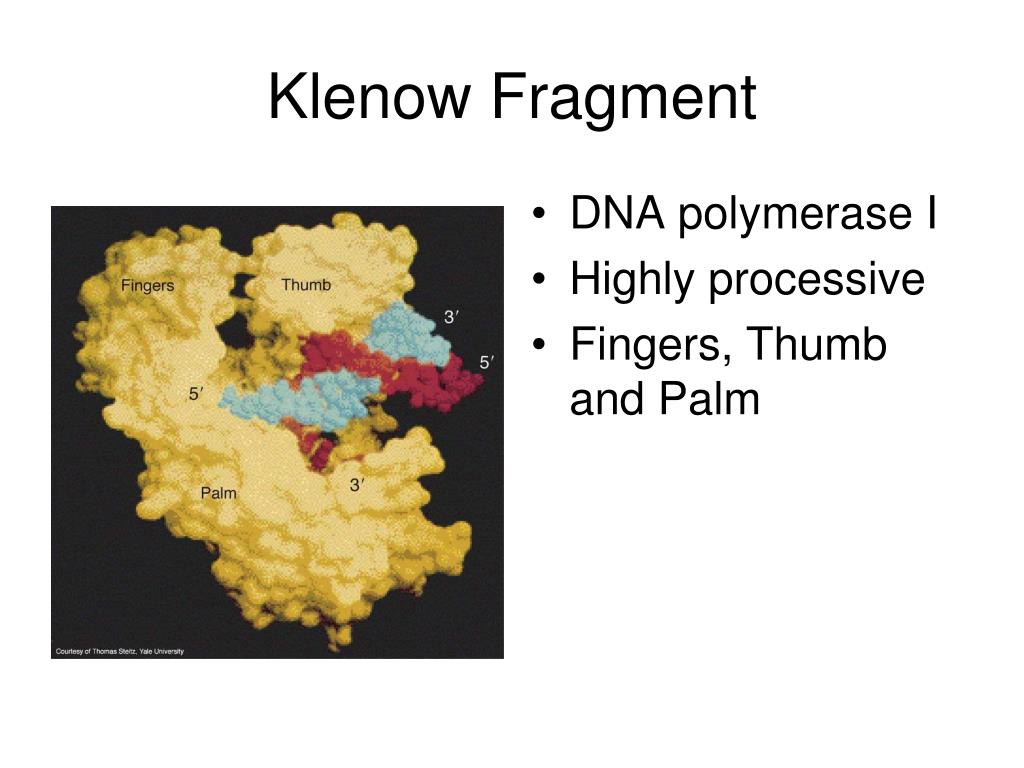
This procedure is frequently used in preparing DNA libraries for next-generation sequencing. 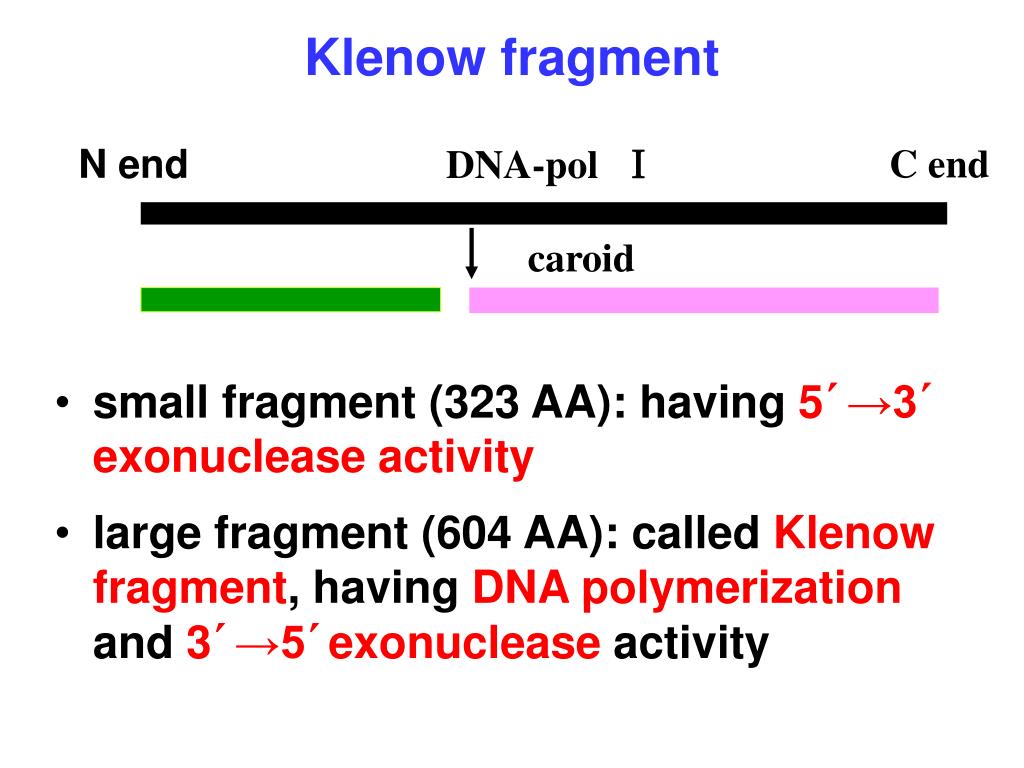
This process of dA-tailing (or dT-tailing) of the DNA is an important step in the process of ligating DNA adapters to DNA fragments. DNA polymerases lacking a 3'→5' exonuclease may add a single nontemplate-dependent nucleotide to the 3'-end of a sequence (tailing) with a preference for dATP depending on the reaction conditions and sequence context. This problem is overcome by introducing mutations that result in the Klenow Fragment (3'→5' exo-) enzyme that retains 5'→3' polymerase activity but lacks any exonuclease activity. coli DNA polymerase I can be undesirable, the 3'→5' exonuclease proofreading activity of Klenow Fragment can also be undesirable in some circumstances. Just as the 5'→3' exonuclease activity of E. coli DNA polymerase I lacks this 5’-exonuclease activity but retains the 5'→3' polymerase activity and the 3'→5' exonuclease activity for removal of precoding nucleotides and proofreading. It is undesirable for DNA sequencing and other applications due to the ability of the 5’→3’ exonuclease to degrade flap structures, single-stranded and double-stranded DNA from free 53'→5'-ends. The 3'→5' nuclease activity is used during DNA repair and is necessary for enzyme applications such as nick translation. The main duty of the 5'→3' exonuclease activity is to remove the RNA primers at the 5' ends of newly synthesized DNA so that the polymerase activity can fill in the resulting gaps. coli DNA polymerase I makes it unsuitable for some applications. The holoenzyme possesses three activities: 5'→3' DNA-dependent DNA polymerase, 3'→5' proofreading and 3'→5' exonuclease (also known as flap endonuclease). DNA polymerase I is a single polypeptide chain consisting of ∼930 residues.
0 Comments
Leave a Reply. |
AuthorWrite something about yourself. No need to be fancy, just an overview. ArchivesCategories |

 RSS Feed
RSS Feed
